Research Insight
Disease Resistance in Canids: Genetic Variations between Wild Wolves and Domestic Dogs 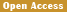
 Author
Author  Correspondence author
Correspondence author
International Journal of Molecular Veterinary Research, 2024, Vol. 14, No. 6 doi: 10.5376/ijmvr.2024.14.0028
Received: 03 Nov., 2024 Accepted: 08 Dec., 2024 Published: 20 Dec., 2024
Lin X.F., 2024, Disease resistance in canids: genetic variations between wild wolves and domestic dogs, International Journal of Molecular Veterinary Research, 14(6): 244-253 (doi: 10.5376/ijmvr.2024.14.0028)
This study explores the comparative genomics of wild wolves and domestic dogs, focusing on genetic differences influencing immunity and disease susceptibility. Using genome-wide association studies (GWAS) and whole-genome sequencing, we identified key genes involved in immune responses and analyzed the effects of domestication on immune gene diversity. Furthermore, evolutionary and ecological factors, including habitat and behavior, were examined for their roles in shaping disease resistance. A case study on rabies resistance in wild wolves highlights adaptive genetic mechanisms and their conservation implications. The findings advance our understanding of canid immunogenomics and suggest strategies for breeding disease-resistant dogs, conserving genetic diversity in wolves, and improving zoonotic disease management. This research bridges genomics with practical applications in veterinary science and wildlife conservation, offering pathways for future exploration of genetic resilience in canids.
1 Introduction
Canids, a family that includes both wild wolves (Canis lupus) and domestic dogs (Canis lupus Familiaris), exhibit significant genetic and phenotypic diversity. Wild wolves are known for their robust health and adaptability to various environments, while domestic dogs have undergone extensive artificial selection, resulting in a wide range of breeds with diverse physical and behavioral traits (Tang et al., 2018; Wang et al., 2018). This domestication process has led to structural variations (SVs) in the genome, which are linked to phenotypic evolution, disease susceptibility, and environmental adaptations. The genetic divergence between wild wolves and domestic dogs provides a unique opportunity to study the impact of domestication on disease resistance and other health-related traits.
Understanding the genetic basis of disease resistance in canids is crucial for several reasons. Firstly, it can provide insights into the evolutionary processes that have shaped the health and survival of these species. For instance, structural variations and copy number variations (CNVs) in the genomes of domestic dogs have been associated with breed-specific disease susceptibilities and phenotypic traits (Marsden et al., 2015; Serres-Armero et al., 2021; Zhang, 2024). Additionally, the domestication process has introduced deleterious genetic variations in dogs due to population bottlenecks and selective breeding practices, which have increased their genetic load compared to wild wolves. Studying these genetic differences can help identify the genes and regulatory elements involved in disease resistance, which is essential for improving breeding programs and conserving genetic diversity in both domestic and wild canids (Ostrander et al., 2019; Mallil et al., 2020).
This study attempts to explore the genetic variations between wild wolves and domestic dogs that contribute to disease resistance, discuss the impact of domestication on genetic load and disease susceptibility in domestic dogs, and provide an overview of the role of epigenetic modifications, such as methylation patterns, in regulating genes associated with disease resistance. Additionally, it aims to offer insights into the evolutionary dynamics of these genetic variations and their implications for the health and conservation of canid species, ultimately contributing to strategies for improving the health and longevity of both wild and domestic populations.
2 Genetic Basis of Disease Resistance in Canids
2.1 Genetic variations and their role in immunity
Genetic variations play a crucial role in the immunity of canids, influencing their ability to resist diseases. Studies have shown that domestication and selective breeding have led to an increase in deleterious genetic variants in domestic dogs compared to their wild counterparts, such as gray wolves. This is primarily due to population bottlenecks and artificial selection, which have reduced the efficiency of natural selection in removing harmful variants (Marsden et al., 2015). Structural variations (SVs) in the genome, including insertions, deletions, and copy number variations (CNVs), also contribute significantly to phenotypic diversity and disease susceptibility in canids. For instance, specific SVs in dogs are enriched in genes related to immune systems, indicating their role in disease resistance (Serres-Armero et al., 2017; Wang et al., 2018; Xuan, 2024).
2.2 Key genes associated with disease resistance
Several key genes have been identified that are associated with disease resistance in canids. For example, the ADGRE1 gene, which is linked to severe malaria resistance in humans, has been found to provide protective host defense against Plasmodium infections in dogs, suggesting its role in resistance to similar parasitic diseases (Figure 1) (Liu et al., 2018). Additionally, genes such as DCK, ICAM4, GAPDH, and BSG, which are related to immune function, are more highly expressed in wolves, indicating a potentially greater resistance to pathogens compared to domestic dogs (Yang et al., 2018). The AKR1B1 gene, which has been duplicated in dogs, is associated with increased antioxidant ability and may contribute to their adaptation to different environments (Wang et al., 2018; Zhao, 2018).
.png) Figure 1 Population structure and genetic diversity of the canids analyzed in this study (Adopted from Liu et al., 2018) Image caption: (A) Geographic locations of the 55 canids studied. (B) Principal component analysis. EGW, Eurasian gray wolves; AGW, African golden wolves; CIDY, Chinese indigenous dogs from Yingjiang; MEVD, Middle Eastern village dogs; EB, European breeds; NID, Nigerian indigenous dogs. (C) Phylogenetic tree using bootstrapping analysis. (D) Structure analysis of the 55 canids. (E) Genetic diversity for the five inferred canid groups (Adopted from Liu et al., 2018) |
2.3 Methods used in genetic analysis
Various methods are employed to analyze genetic variations and their implications for disease resistance in canids. Whole genome sequencing (WGS) is a powerful tool that provides comprehensive insights into the genetic makeup of both domestic dogs and wild canids. It has been used to identify genome-wide patterns of deleterious variation and to create detailed maps of structural variations (Marsden et al., 2015; Serres-Armero et al., 2017). Genome-wide association studies (GWAS) are another critical method, particularly useful for identifying loci associated with specific traits and disease susceptibilities. For example, a CNV-based GWAS identified 96 loci with copy number differences across dog breeds, which are associated with breed-specific morphometrics and disease susceptibilities (Serres-Armero et al., 2021). Additionally, transcriptome analysis using RNA-Seq technology helps in understanding gene expression differences between dogs and wolves, providing insights into their immune responses and other physiological traits (Koch et al., 2016; Yang et al., 2018).
Genetic variations, including single nucleotide variants, structural variations, and copy number variations, play a significant role in the disease resistance of canids. Key genes such as ADGRE1 and AKR1B1 have been identified as crucial for immune function and adaptation. Advanced genetic analysis methods like Whole Genome Sequencing and Genome-Wide Association Studies are essential tools for uncovering the genetic basis of disease resistance in canids. These findings highlight the complex interplay between genetics and disease resistance, influenced by domestication and selective breeding practices.
3 Comparative Analysis of Wild Wolves and Domestic Dogs
3.1 Evolutionary pathways and genetic divergence
The evolutionary pathways of wild wolves and domestic dogs have diverged significantly due to domestication. Structural variations (SVs) in the genome have played a crucial role in this divergence, influencing phenotypic evolution, disease susceptibility, and environmental adaptations. For instance, the dog genome has accumulated specific insertions, deletions, and repeats that are not present in wolves, such as a novel copy of the AKR1B1 gene, which is involved in fatty acid synthesis and antioxidant ability, likely in response to dietary shifts during domestication (Wang et al., 2018; Wei, 2018). Additionally, copy number variations (CNVs) have been identified in both dogs and wolves, with significant differences in loci responsible for sensory perception, immune response, and metabolic processes (Figure 2) (Serres-Armero et al., 2017; Serres-Armero et al., 2021).
.png) Figure 2 Landscape of canine segmental duplications (Adopted from Serres-Armero et al., 2017) Image caption: a Genome-wide map of canine SDs. Autosomes are represented by horizontal bars, and each mark represent a duplicated region identified in at least one sample of the group indicated. b Total length of genomic duplications identified per subspecies. c Venn diagram showing intersection of duplicated regions identified in dogs, gray wolves and canines in the outgroup (chrUn excluded) (Adopted from Serres-Armero et al., 2017) |
3.2 Impact of domestication on immune gene diversity
Domestication has had a profound impact on the immune gene diversity of dogs compared to their wild counterparts. The process of domestication, characterized by population bottlenecks and selective breeding, has increased the number of deleterious genetic variants in dogs. This is evident in the higher genetic load observed in domestic dogs compared to gray wolves, which is attributed to less efficient natural selection during domestication and breed formation (Marsden et al., 2015). Furthermore, studies on Toll-like receptors (TLRs) have shown that domestic dogs possess protective alleles against inflammatory bowel disease (IBD) that are absent in wild canids like the maned wolf and red wolf, suggesting that these protective alleles developed post-domestication (Henson et al., 2017).
3.3 Overview of disease profiles in wolves vs dogs
The disease profiles of wild wolves and domestic dogs differ significantly due to genetic and environmental factors. Domestic dogs have been found to carry a higher burden of deleterious genetic variants, which can lead to a variety of breed-specific diseases (Marsden et al., 2015). For example, African indigenous dogs have developed genetic adaptations to tropical parasites, including genes linked to immunity and resistance to severe malaria, which are not present in wild wolves (Liu et al., 2018). In contrast, wild wolves, such as those in Yellowstone National Park, show a relationship between reduced genomic variation and increased severity of diseases like sarcoptic mange, highlighting the importance of genetic diversity in disease resistance (DeCandia et al., 2020). Additionally, the lack of protective alleles against IBD in wild canids further underscores the differences in disease susceptibility between wild and domestic canids (Henson et al., 2017).
The comparative analysis of wild wolves and domestic dogs reveals significant genetic divergence due to domestication, impacting immune gene diversity and disease profiles. Structural and copy number variations have played a crucial role in this divergence, with domestic dogs showing higher genetic loads and specific adaptations to their environments. The differences in disease susceptibility and genetic diversity between wild and domestic canids highlight the complex interplay between genetics and domestication in shaping the health and evolution of these species.
4 Immune System Differences in Canids
4.1 Innate immunity: genetic and molecular insights
Innate immunity, the first line of defense against pathogens, shows significant genetic variation between wild wolves and domestic dogs. Studies have identified several genes related to the innate immune response that are differentially expressed between these canids. For instance, genes such as CCL23, TRIM10, DUSP10, RAB27A, CLEC5A, and GCH1 are more highly expressed in wolves, suggesting a stronger innate immune response compared to dogs (Yang et al., 2018). Additionally, Toll-like receptors (TLRs), which play a crucial role in pathogen recognition, exhibit notable polymorphisms in wild canids. For example, the TLR5 gene in red wolves and maned wolves shows novel polymorphic positions not found in domestic dogs, indicating potential differences in disease susceptibility and immune response (Henson et al., 2017). These genetic variations highlight the evolutionary adaptations in the innate immune system of wild canids, possibly due to their exposure to a broader range of pathogens in the wild.
4.2 Adaptive immunity: diversity in t-cell receptor and mhc complexes
The adaptive immune system, particularly the diversity in T-cell receptors (TCRs) and major histocompatibility complex (MHC) genes, also differs significantly between wild wolves and domestic dogs. MHC genes, which are critical for antigen presentation and immune response, show varying levels of polymorphism. In wild canids, such as the European roe deer, high levels of amino acid diversity in MHC genes have been observed, suggesting a robust adaptive immune response to diverse pathogens (Quéméré et al., 2015). In contrast, domestic dogs have undergone genetic bottlenecks and selective breeding, leading to reduced MHC diversity and potentially higher susceptibility to certain diseases (Marsden et al., 2015). This reduced genetic diversity in domestic dogs' MHC complexes may compromise their ability to respond to new or evolving pathogens effectively.
4.3 Genetic bottlenecks and consequences for disease susceptibility
The domestication of dogs has involved significant genetic bottlenecks and selective sweeps, which have increased the prevalence of deleterious genetic variants. These bottlenecks have led to a higher genetic load in domestic dogs compared to wild wolves, with dogs exhibiting 2-3% higher genetic load on average (Marsden et al., 2015). This increased genetic load is associated with a higher incidence of breed-specific diseases and reduced overall fitness. For example, the lack of protective alleles in the TLR5 gene in domestic dogs, which are present in some wild canids, suggests a higher predisposition to inflammatory diseases such as inflammatory bowel disease (IBD) (Henson et al., 2017). Additionally, the enrichment of copy number variations (CNVs) in genes related to immune response in domestic dogs further underscores the impact of domestication on disease susceptibility (Serres-Armero et al., 2017; Kessler et al., 2020). These genetic changes highlight the trade-offs associated with domestication, where artificial selection for specific traits has inadvertently increased the risk of certain diseases in domestic dogs.
The immune system differences between wild wolves and domestic dogs are marked by significant genetic variations in both innate and adaptive immunity. Wild canids exhibit greater genetic diversity in immune-related genes, which likely contributes to their robust immune responses. In contrast, domestic dogs have experienced genetic bottlenecks and selective breeding, leading to reduced genetic diversity and increased susceptibility to certain diseases. These findings underscore the complex interplay between domestication, genetic variation, and disease resistance in canids.
5 Environmental and Behavioral Influences on Disease Resistance
5.1 Influence of habitat and lifestyle on disease exposure
The habitat and lifestyle of canids significantly influence their exposure to diseases. Wild wolves, for instance, inhabit diverse and often harsh environments that expose them to a wide range of pathogens. This constant exposure can lead to a more robust immune system due to natural selection favoring individuals with better disease resistance. In contrast, domestic dogs often live in more controlled environments with less pathogen exposure, which can result in a less diverse immune response (Figure 3) (Marsden et al., 2015; Pilot et al., 2021).
.png) Figure 3 Distribution of samples of Eurasian grey wolves (red circles) and free-ranging dogs (green circles) analysed in this study (Adopted from Pilot et al., 2021) Image caption: Geographic locations of samples are precise except Mongolian wolves and free-ranging dogs from China and Portugal, which have approximate locations. The number of samples collected from the same locations is reflected by the circle size. The introgression pattern analysis was carried out for West Eurasian wolves (marked with the red frame) and all free-ranging dogs shown on the map that carried introgressed chromosomal fragments. Among East Asian wolves (black frame), only four admixed individuals were found, including an F1 hybrid (Adopted from Pilot et al., 2021) |
Domestication has also led to population bottlenecks and selective breeding, which have increased the genetic load of deleterious variants in domestic dogs compared to wild wolves. This increased genetic load can make domestic dogs more susceptible to certain diseases (Marsden et al., 2015). Additionally, the shift in diet and lifestyle during domestication has led to structural variations in the genome of domestic dogs, which are associated with changes in immune system function and disease susceptibility (Wang et al, 2018).
5.2 Interaction between behavior and immune responses
Behavioral factors play a crucial role in the immune responses of canids. For example, the social structure and pack behavior of wild wolves can influence disease transmission and immune system development. Wolves living in packs may have a higher exposure to pathogens due to close contact with other pack members, which can lead to a more robust immune response (DeCandia et al., 2020).
In domestic dogs, selective breeding for specific behaviors and traits has also impacted their immune responses. For instance, certain breeds have been found to have higher incidences of specific diseases due to the genetic bottlenecks and inbreeding associated with breed formation. Moreover, the interaction between behavior and immune response is evident in the case of inflammatory bowel disease (IBD) in canids. Studies have shown that domestic dogs have developed protective alleles against IBD, which are not present in wild canids like the red wolf and maned wolf, suggesting that these protective alleles may have developed post-domestication due to changes in behavior and lifestyle (Henson et al., 2017).
In summary, the environmental and behavioral factors significantly influence disease resistance in canids. Wild wolves, with their diverse habitats and social behaviors, tend to have more robust immune systems compared to domestic dogs, which have undergone genetic changes due to domestication and selective breeding. These differences highlight the complex interplay between environment, behavior, and genetics in shaping disease resistance in canids.
6 Case Study: Rabies Resistance in Wild Wolves
6.1 Genetic and epidemiological insights into rabies resistance
Rabies is a significant viral disease affecting both domestic and wild canids. In wild wolves, genetic variations and epidemiological factors play crucial roles in their resistance to rabies. Studies have shown that wild wolves, like other wild carnivores, can be reservoirs for the rabies virus (RABV). For instance, an outbreak in the Madikwe Game Reserve in South Africa highlighted the presence of canid RABVs in wild dogs and other carnivores, suggesting a complex interaction between wild and domestic animals in the transmission dynamics of rabies (Sabeta et al., 2018). Genetic analyses of these viruses revealed phylogenetic clustering with RABVs from domestic dogs and other wild carnivores, indicating potential cross-species transmission and the importance of genetic monitoring in understanding rabies resistance.
6.2 Adaptive mechanisms in wolves compared to domestic dogs
The genetic makeup of wild wolves has been shaped by natural selection, which may confer certain advantages in disease resistance compared to domestic dogs. Structural variations (SVs) in the genomes of wolves and dogs have been linked to differences in immune system function and disease susceptibility. For example, studies have identified specific SVs in dogs that are associated with immune response genes, which may differ from those in wolves due to the domestication process (Serres-Armero et al., 2017; Wang et al., 2018). Additionally, the presence of endogenous retroviruses (ERVs) in the genomes of canids, including wolves, suggests a historical adaptation to viral infections, which could influence their current resistance to diseases like rabies (Halo et al., 2019).
6.3 Implications for conservation and disease management
Understanding the genetic and adaptive mechanisms underlying rabies resistance in wild wolves has significant implications for conservation and disease management. Conservation strategies should consider the genetic diversity and adaptive potential of wild wolf populations to ensure their resilience against diseases. Regular vaccination programs for at-risk wildlife, as well as monitoring and controlling rabies in domestic animals, are crucial to prevent outbreaks and protect endangered species (Sabeta et al., 2018). Moreover, maintaining large population sizes and genetic diversity in wild wolves can help mitigate the effects of genetic drift and inbreeding, which are critical for the long-term survival and health of these populations (Marsden et al., 2015; Pilot et al., 2021).
The study of rabies resistance in wild wolves reveals important genetic and epidemiological insights that differentiate them from domestic dogs. Adaptive mechanisms, shaped by natural selection, provide wild wolves with certain advantages in disease resistance. These findings underscore the importance of genetic diversity and proactive disease management in conservation efforts to protect wild canid populations.
7 Applications of Genetic Findings
7.1 Breeding strategies for enhanced disease resistance in dogs
Genetic research has revealed that domestication and selective breeding have led to an increased burden of deleterious genetic variants in domestic dogs compared to their wild counterparts, such as gray wolves. This is primarily due to population bottlenecks and intense artificial selection for breed-specific traits, which have reduced the efficiency of natural selection in removing harmful variants (Marsden et al., 2015). To enhance disease resistance in dogs, breeding strategies should focus on maintaining larger population sizes and promoting genetic diversity. Avoiding the overly typological practice of breeding individuals that best fit breed standards can help reduce the accumulation of deleterious variants. Additionally, leveraging resources like iDog, which integrates genomic data and disease traits, can aid in identifying and selecting for genetic markers associated with disease resistance (Tang et al., 2018).
7.2 Contributions to conservation genetics of wild wolves
The genetic insights gained from studying domestic dogs can also be applied to the conservation genetics of wild wolves. For instance, understanding the impact of structural variations (SVs) and copy number variations (CNVs) on phenotypic traits and disease susceptibility in dogs can inform conservation strategies for wolves. Research has shown that SVs and CNVs play significant roles in immune response and metabolic processes, which are crucial for the survival of wild populations (Wang et al., 2018; Serres-Armero et al., 2021). By identifying and preserving genetic diversity in wild wolf populations, conservation efforts can mitigate the risks associated with small population sizes and inbreeding, which are known to increase the genetic load of deleterious variants (Marsden et al., 2015; Serres-Armero et al., 2017).
7.3 Insights for zoonotic disease control
The genetic relationship between domestic dogs and wild canids, such as wolves, provides valuable insights for zoonotic disease control. Hybridization between domestic dogs and wild canids, including African wolves, has been documented, leading to gene flow and potential disease transmission between species (Mallil et al., 2020). Understanding the genetic basis of disease resistance and susceptibility in both domestic and wild canids can help in developing strategies to control zoonotic diseases. For example, identifying genes associated with immune response and disease resistance in dogs can inform vaccination and disease management programs for both domestic and wild canid populations (Wang et al., 2018; Serres-Armero et al., 2021). Additionally, maintaining genetic diversity and large population sizes in wild canids can reduce the prevalence of disease and the risk of zoonotic transmission (Marsden et al., 2015; Pilot et al., 2021). Genetic research on domestic dogs and wild wolves offers significant applications in breeding strategies, conservation genetics, and zoonotic disease control. By promoting genetic diversity and understanding the genetic basis of disease resistance, we can enhance the health and survival of both domestic and wild canid populations.
8 Challenges and Future Directions
8.1 Knowledge gaps in canid immunogenomics
Despite significant advancements in canid genomics, there remain substantial gaps in our understanding of the immunogenomic differences between wild wolves and domestic dogs. Structural variations (SVs) and copy number variations (CNVs) have been identified as key factors influencing disease susceptibility and immune responses in canids. For instance, SVs in dogs are enriched in genes associated with immune systems, suggesting a potential role in disease resistance and susceptibility (Wang et al., 2018). However, comprehensive genome-wide studies linking these variations to specific immune responses are still lacking. Additionally, the increased genetic load in domestic dogs due to population bottlenecks and selective breeding further complicates our understanding of their immune capabilities compared to wild wolves (Marsden et al., 2015).
8.2 Integrating genomics with conservation and veterinary practices
The integration of genomic data into conservation and veterinary practices holds promise for improving the health and survival of both wild and domestic canids. Resources like iDog, which consolidate genomic, phenotypic, and disease-related data, are invaluable for researchers and practitioners (Tang et al., 2018). These tools can aid in identifying genetic markers for disease resistance and inform breeding programs to reduce the prevalence of deleterious genetic variants. Moreover, understanding the genomic basis of disease susceptibility can help in developing targeted veterinary treatments and conservation strategies. For example, the identification of CNVs associated with breed-specific disease susceptibilities can guide selective breeding practices to enhance genetic diversity and reduce the incidence of hereditary diseases (Serres-Armero et al., 2021).
8.3 Prospects for CRISPR and other gene-editing tools
The advent of CRISPR and other gene-editing technologies offers exciting prospects for addressing genetic diseases in canids. CRISPR systems have already shown potential in tracking and controlling the spread of antimicrobial resistance genes in bacteria of canine origin, highlighting their utility in managing infectious diseases (Rossi et al., 2019). In the context of canid genomics, CRISPR could be employed to correct deleterious genetic variants identified through genome-wide association studies (GWAS) and other genomic analyses. This approach could mitigate the genetic load in domestic dogs and enhance their disease resistance. However, the ethical and practical implications of gene editing in wild populations must be carefully considered to avoid unintended ecological consequences.
Addressing the challenges in canid immunogenomics requires filling knowledge gaps through comprehensive genomic studies, integrating genomic data into conservation and veterinary practices, and exploring the potential of gene-editing technologies like CRISPR. These efforts can lead to improved health outcomes for both wild wolves and domestic dogs, ensuring their survival and well-being in a rapidly changing world.
9 Concluding Remarks
The study of genetic variations between wild wolves and domestic dogs has revealed significant insights into the evolutionary dynamics and disease resistance in canids. Structural variations (SVs) and copy number variations (CNVs) have been identified as key factors influencing phenotypic evolution, disease susceptibility, and environmental adaptations in domestic dogs compared to their wild counterparts. For instance, the presence of a new AKR1B1 gene copy in domestic dogs, which is associated with increased fatty acid synthesis and antioxidant ability, highlights the impact of dietary shifts during domestication. Additionally, the domestication process has led to an increased burden of deleterious genetic variants in dogs due to population bottlenecks and artificial selection, which has implications for breed-specific traits and disease susceptibility. The integration of genomic data from various canid species through resources like iDog has further enriched our understanding of the genetic landscape and its functional relevance.
The findings from this study have profound implications for both veterinary and conservation sciences. In veterinary science, understanding the genetic basis of disease susceptibility in domestic dogs can inform selective breeding programs aimed at reducing the prevalence of deleterious variants and improving overall canine health. The identification of specific CNVs and SVs associated with disease traits can lead to the development of targeted genetic tests and personalized treatment plans for dogs. In conservation science, the evidence of hybridization between wild canids, such as the African wolf, and domestic dogs raises concerns about genetic dilution and the potential loss of unique genetic lineages. Conservation strategies must therefore consider the impact of gene flow from domestic dogs on wild canid populations and implement measures to preserve genetic diversity and prevent disease transmission.
Bridging the gap between research and practical application is crucial for maximizing the benefits of genetic studies in canids. The comprehensive genomic data and resources developed through this research provide a valuable foundation for future studies and applications. For instance, the iDog database serves as an integrated platform for accessing and analyzing genetic information, which can facilitate collaboration among researchers, veterinarians, and conservationists. By leveraging these resources, stakeholders can develop more effective breeding programs, conservation strategies, and disease management plans that are informed by the latest genetic insights. Ultimately, the integration of research findings into practical applications will enhance the health and sustainability of both domestic and wild canid populations.
Acknowledgments
I am grateful to Dr. Luo for critically reading the manuscript and providing valuable feedback that improved the clarity of the text. We express our heartfelt gratitude to the two anonymous reviewers for their valuable comments on the manuscript.
Conflict of Interest Disclosure
The author affirms that this research was conducted without any commercial or financial relationships that could be construed as a potential conflict of interest.
DeCandia A., Schrom E., Brandell E., Stahler D., and Vonholdt B., 2020, Sarcoptic mange severity is associated with reduced genomic variation and evidence of selection in Yellowstone National Park wolves (Canis lupus), Evolutionary Applications, 14: 429-445.
https://doi.org/10.1111/eva.13127
Halo J., Pendleton A., Jarosz A., Gifford R., Day M., and Kidd J., 2019, Origin and recent expansion of an endogenous gammaretroviral lineage in domestic and wild canids, Retrovirology, 16: 1-25.
https://doi.org/10.1186/s12977-019-0468-z
Henson L., Songsasen N., Waddell W., Wolf K., Emmons L., González S., Freeman E., and Maldonado J., 2017, Characterization of genetic variation and basis of inflammatory bowel disease in the toll-like receptor 5 gene of the red wolf and maned wolf, Endangered Species Research, 32: 135-144.
https://doi.org/10.3354/ESR00790
Kessler C., Brambilla A., Waldvogel D., Camenisch G., Biebach I., Leigh D., Grossen, C., and Croll D., 2020, A robust sequencing assay of a thousand amplicons for the high‐throughput population monitoring of Alpine ibex immunogenetics, Molecular Ecology Resources, 22: 66-85.
https://doi.org/10.1111/1755-0998.13452
Koch I., Clark M., Thompson M., Deere-Machemer K., Wang J., Duarte L., Gnanadesikan G., McCoy E., Rubbi L., Stahler D., Pellegrini M., Ostrander E., Wayne R., Sinsheimer J., and Vonholdt B., 2016, The concerted impact of domestication and transposon insertions on methylation patterns between dogs and grey wolves, Molecular Ecology, 25(8): 1838-1855.
https://doi.org/10.1111/mec.13480
Liu Y., Wang L., Xu T., Guo X., Li Y., Yin T., Yang H., Hu Y., Adeola A., Sanke O., Otecko N., Wang M., Ma Y., Charles O., Sinding M., Gopalakrishnan S., Samaniego J., Hansen A., Fernandes C., Gaubert P., Budd J., Dawuda P., Rueness, E., Jiang L., Zhai W., Gilbert T., Peng M., Qi X., Wang G., and Zhang Y., 2018, Whole‐genome sequencing of African dogs provides insights into adaptations against tropical parasites, Molecular Biology and Evolution, 35: 287-298.
https://doi.org/10.1093/molbev/msx258
Mallil K., Justy F., Rueness E., Dufour S., Totis T., Bloch C., Baarman J., Amroun M., and Gaubert P., 2020, Population genetics of the African wolf (Canis lupaster) across its range: first evidence of hybridization with domestic dogs in Africa, Mammalian Biology, 100: 645-658.
https://doi.org/10.1007/s42991-020-00059-1
Marsden C., Vecchyo D., O’Brien D., Taylor J., Ramírez O., Vilà C., Marquès-Bonet T., Schnabel R., Wayne R., and Lohmueller K., 2015, Bottlenecks and selective sweeps during domestication have increased deleterious genetic variation in dogs, Proceedings of the National Academy of Sciences, 113: 152-157.
https://doi.org/10.1073/pnas.1512501113
Ostrander E., Wang G., Larson G., Vonholdt B., Davis B., Jagannathan V., Hitte C., Wayne R., and Zhang Y., 2019, Dog10K: an international sequencing effort to advance studies of canine domestication, phenotypes and health, National Science Review, 6: 810-824.
https://doi.org/10.1093/nsr/nwz049
Pilot M., Moura A., Okhlopkov I., Mamaev N., Manaseryan N., Hayrapetyan V., Kopaliani N., Tsingarska E., Alagaili A., Mohammed O., Ostrander E., and Bogdanowicz W., 2021, Human‐modified canids in human‐modified landscapes: The evolutionary consequences of hybridization for grey wolves and free‐ranging domestic dogs, Evolutionary Applications, 14: 2433-2456.
https://doi.org/10.1111/eva.13257
Quéméré E., Galan M., Cosson J., Klein F., Aulagnier S., Gilot‐Fromont E., Merlet J., Bonhomme M., Hewison A., and Charbonnel N., 2015, Immunogenetic heterogeneity in a widespread ungulate: the European roe deer (Capreolus capreolus), Molecular Ecology, 24(15): 3873-3887.
https://doi.org/10.1111/mec.13292
Rossi C., Andrade-Oliveira A., and Giambiagi-deMarval M., 2019, CRISPR tracking reveals global spreading of antimicrobial resistance genes by Staphylococcus of canine origin, Veterinary Microbiology, 232: 65-69.
https://doi.org/10.1016/J.VETMIC.2019.04.009
Sabeta C., Rensburg D., Phahladira B., Mohale D., Harrison-White R., Esterhuyzen C., and Williams J., 2018, Rabies of canid biotype in wild dog (Lycaon pictus) and spotted hyaena (Crocuta crocuta) in Madikwe Game Reserve, South Africa in 2014-2015: diagnosis, possible origins and implications for control, Journal of the South African Veterinary Association, 89(1): 1-13.
https://doi.org/10.4102/jsava.v89i0.1517
Serres-Armero A., Davis B., Povolotskaya I., Morcillo-Suarez C., Plassais J., Juan D., Ostrander E., and Marquès-Bonet T., 2021, Copy number variation underlies complex phenotypes in domestic dog breeds and other canids, Genome Research, 31: 762-774.
https://doi.org/10.1101/gr.266049.120
Serres-Armero A., Povolotskaya I., Quilez J., Ramírez O., Santpere G., Kuderna L., Hernandez-Rodriguez J., Fernández-Callejo M., Gómez-Sánchez D., Freedman A., Fan Z., Novembre J., Navarro A., Boyko A., Wayne R., Vilà C., Lorente-Galdos B., and Marquès-Bonet T., 2017, Similar genomic proportions of copy number variation within gray wolves and modern dog breeds inferred from whole genome sequencing, BMC Genomics, 18: 1-15.
https://doi.org/10.1186/s12864-017-4318-x
Tang B., Zhou Q., Dong L., Li W., Zhang X., Lan L., Zhai S., Xiao J., Zhang Z., Bào Y., Zhang Y., Wang G., and Zhao W., 2018, iDog: an integrated resource for domestic dogs and wild canids, Nucleic Acids Research, 47: D793-D800.
https://doi.org/10.1093/nar/gky1041
Wang G., Shao X., Bai B., Wang J., Wang X., Cao X., Liu Y., Wang X., Yin T., Zhang S., Lu Y., Wang Z., Wang L., Zhao W., Zhang B., Ruan J., and Zhang Y., 2018, Structural variation during dog domestication: insights from gray wolf and dhole genomes, National Science Review, 6: 110-122.
https://doi.org/10.1093/nsr/nwy076
Wei F., 2018, Structural variation provides novel insights into dog domestication, National Science Review, 6: 123.
https://doi.org/10.1093/nsr/nwy099
Xuan J., 2024, The genetic basis of flocking behavior in sheep: discoveries from genome-wide association studies, Animal Molecular Breeding, 14(1): 86-94.
https://doi.org/10.5376/amb.2024.14.0011
Yang X., Zhang H., Shang J., Liu G., Xia T., Zhao C., Sun G., and Dou H., 2018, Comparative analysis of the blood transcriptomes between wolves and dogs, Animal Genetics, 49: 291-302.
https://doi.org/10.1111/age.12675
Zhao F., 2018, Mining the hidden treasures from canid genomes, National Science Review, 6: 124.
https://doi.org/10.1093/nsr/nwy087
Zhang A.T., 2024, Research on the role of DNA methylation in the epigenetic regulation mechanism of Pomeranian dogs, Animal Molecular Breeding, 14(1): 72-81.
https://doi.org/10.5376/amb.2024.14.0009

. PDF(466KB)
. FPDF(win)
. FPDF(mac)
. HTML
. Online fPDF
Associated material
. Readers' comments
Other articles by authors
. Xiaofang Lin
Related articles
. Canid genomics
. Disease resistance
. Wild wolves
. Domestic dogs
. Immune gene diversity
Tools
. Email to a friend
. Post a comment


